Increasing the means of industrial education and extending the influence of science and art upon productive industry
Award Categories
About Us
MoreIncreasing the means of industrial education and extending the influence of science and art upon productive industry.
Awards
MoreThe Royal Commission for the Exhibition of 1851 awards some 35 postgraduate Fellowships and Scholarships a year, for advanced study and research in science, engineering, design and the built environment.
Special Awards
MOREA very limited number of Special Awards are made to worthy causes and individuals whose aims are consistent with the Commission’s Charter ‘to increase the means of industrial education’ in Britain. These may range from substantial support for other bodies in pursuing specific projects, to travel and study awards to individuals.
History
& Archive
More
The Archive of the Royal Commission for the Exhibition 1851 dates from 1849 when the Society of Arts and its president Prince Albert, the Prince Consort, conceived the idea of holding a Great Exhibition in London in 1851.

News & Events
More-

Royal Commission boosts UK Research & Development with Fellowships linking academia and industry
-

-

-

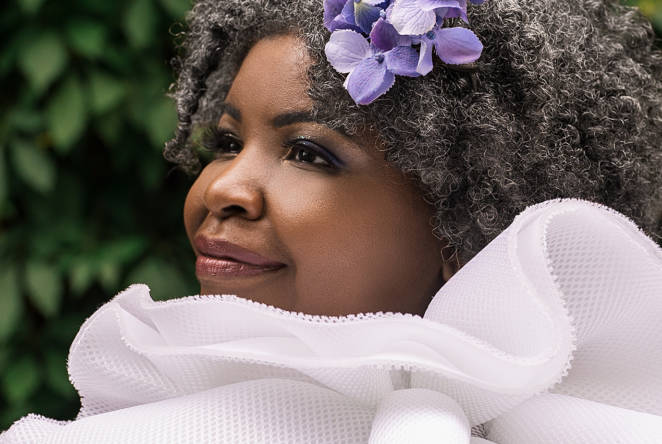
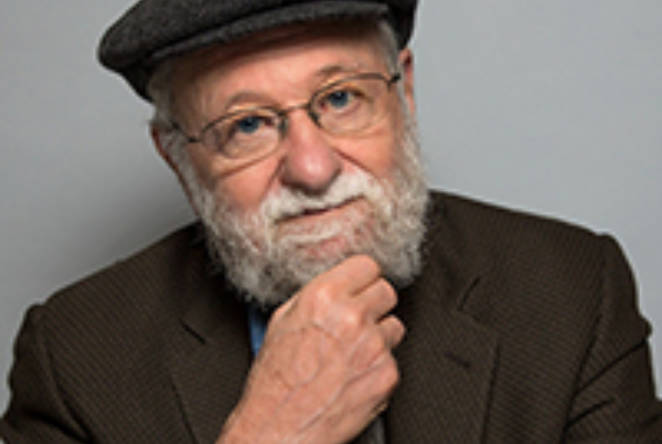
2023 Sir Misha Black Award for Innovation in Design Education to be given to Haleh Moravej
-

-

-

-
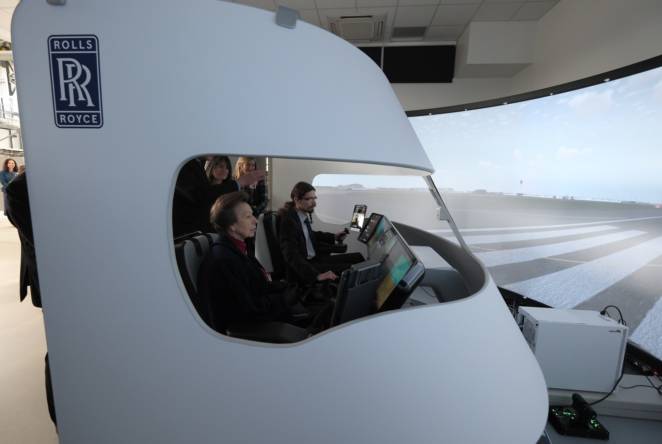

The Princess Royal officially opens Cranfield University's flying classroom
-
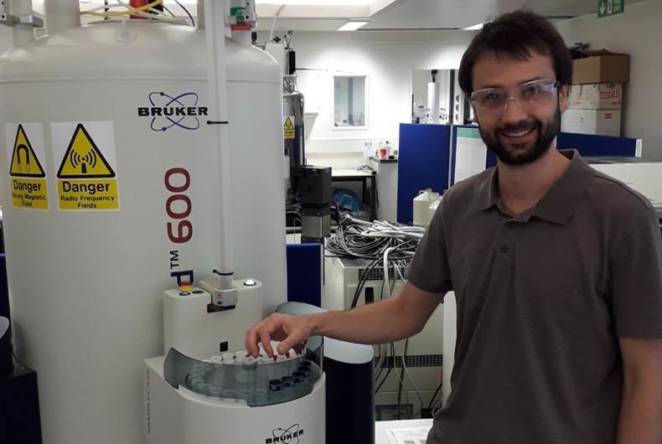

Research Fellow's work now out in Chemistry- A European Journal
-
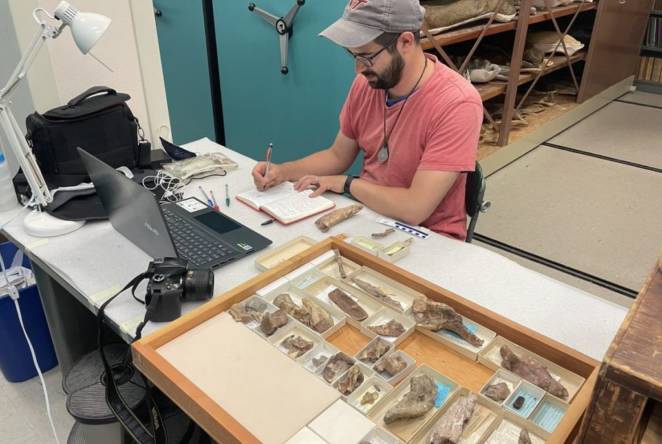

-

Royal Commission announces first Regenerative Design Fellows
-
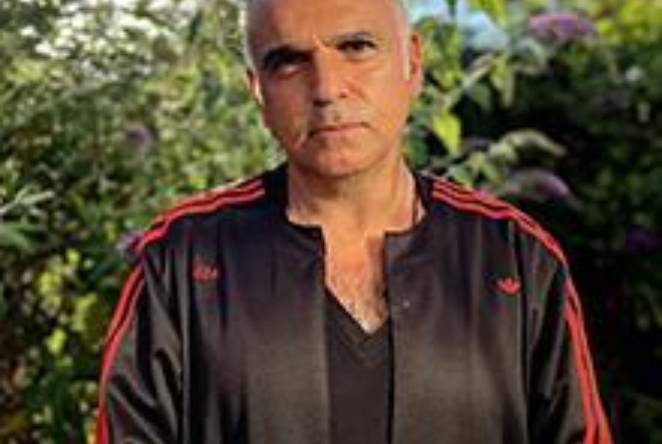

2022 Sir Misha Black Award for Innovation in Design Education to be given to Judah Armani
-

-
-
-

-

-
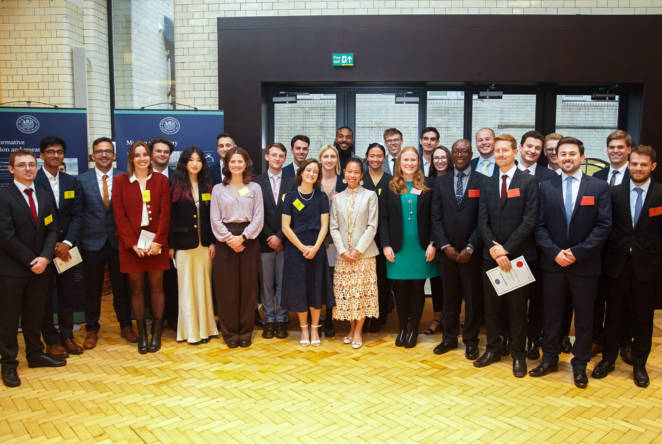

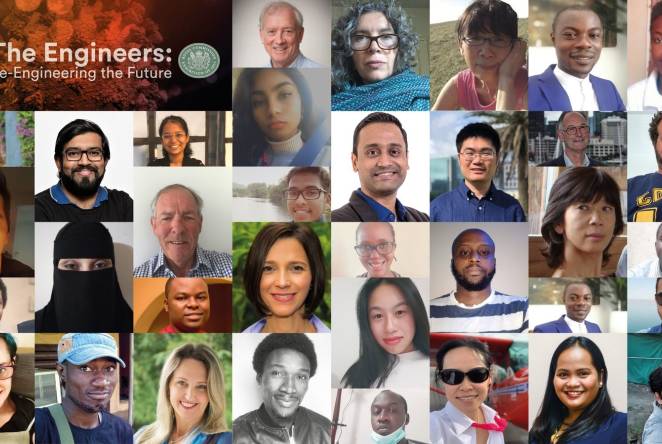
Royal Commission backs UK Industry and innovation to invigorate British research and development
-
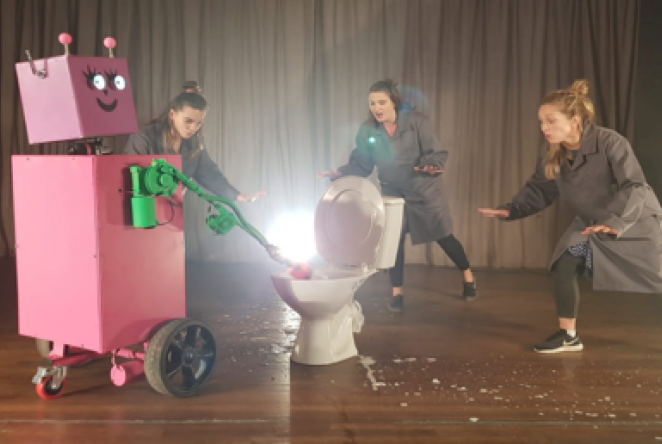

1851 Special Awardee, Kids Invent Stuff, create an original engineering-themed musical
-

1851 Royal Commission announces new fellowship in Regenerative Design
-
-
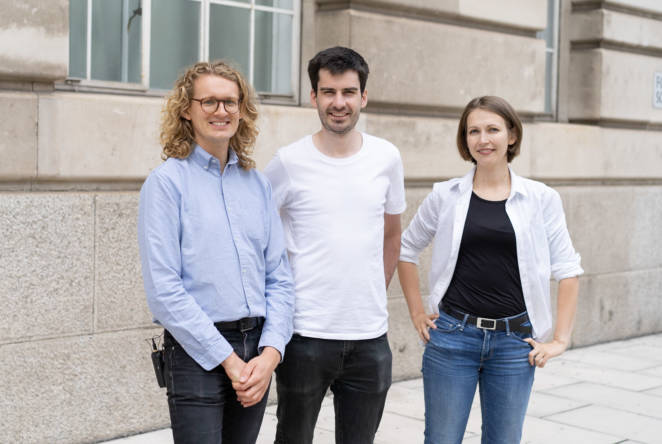

RAEng 1851 Enterprise Fellow building the next generation of fire-safe insulation products
-

-
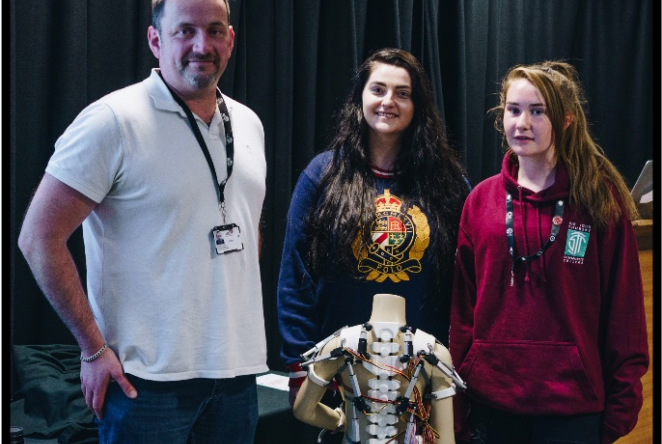

1851 Special Awardee, Primary Engineer, releases new podcast series: If You Were An Engineer"
-

-
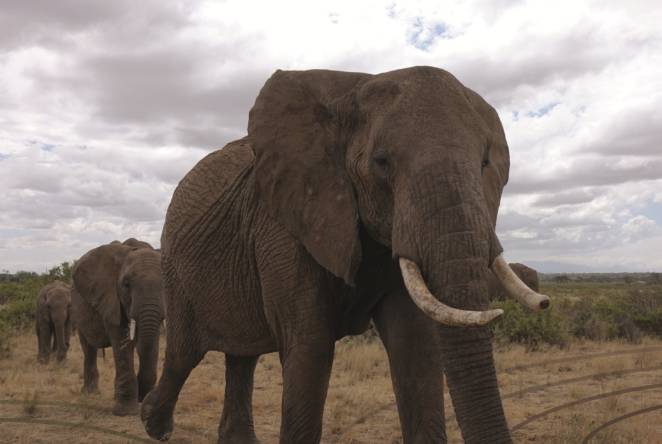

1851 Alumna publishes paper in collaboration with 'Save The Elephants'
-
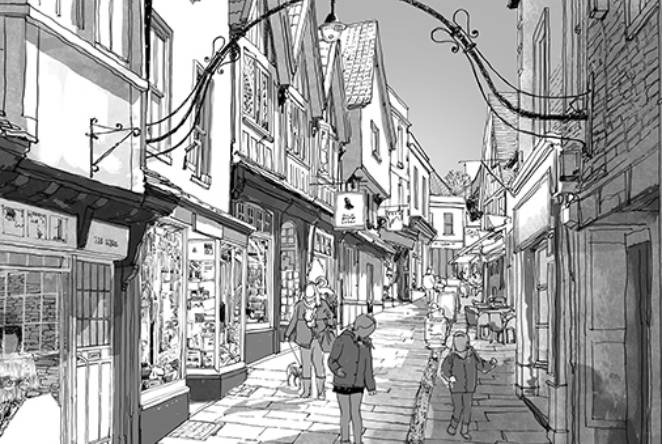

From recessions to resurgence: the Fellowship revealing the secrets of the high street
-
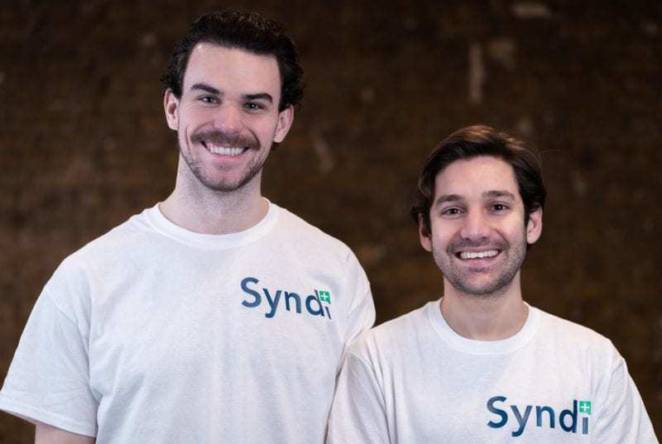

-
Meet the 1851 Fellows making a difference in the UK's pandemic response
-
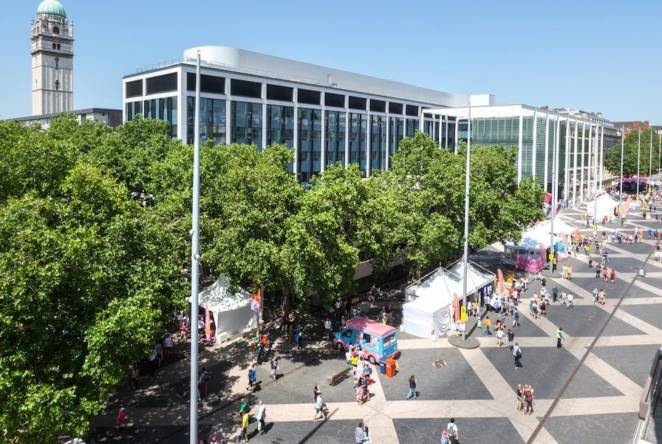

-
-
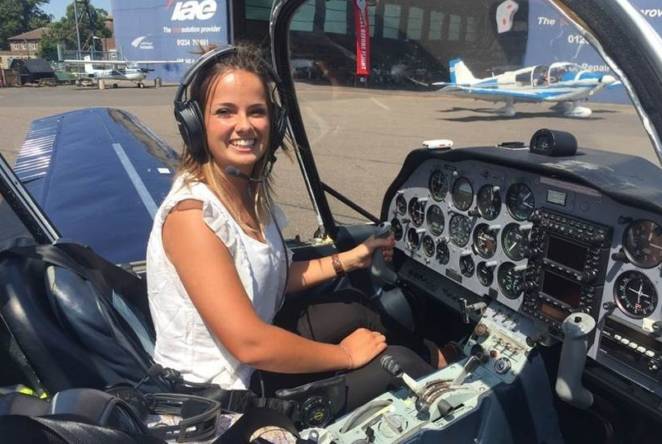

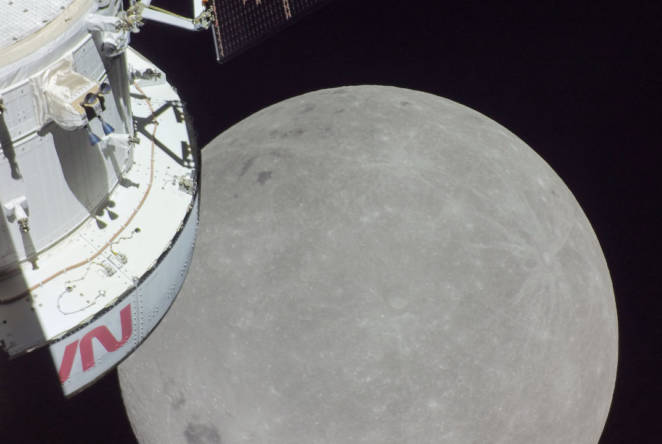
After NASA touchdown, meet the 1851 Industrial Fellow on a mission to Mars
-

RAEng 1851 Enterprise Fellow receives Award from UK Gov for ‘Innovative’ and ‘Inspiring’ Project
-

-
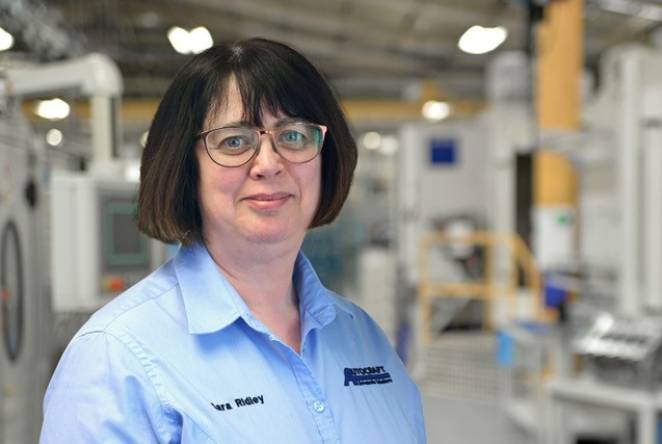

Celebrating IWD with Dr Sara Ridley, Industrial Fellow and STEM ambassador
-
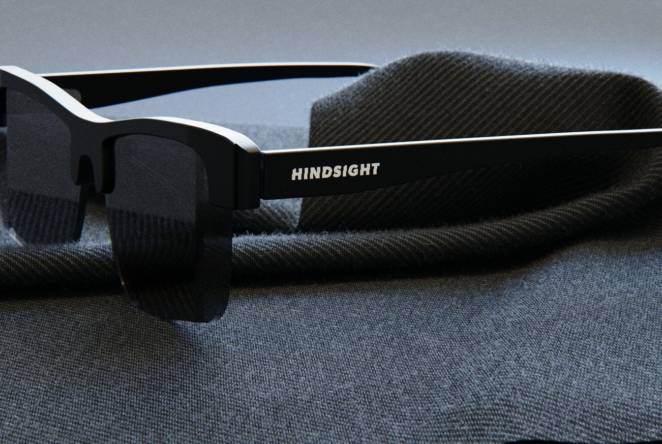

RAEng 1851 Enterprise Fellow launches revolutionary new glasses
-

-

The Commission celebrates work of Fellows with Alumni Science Evening
-
-

-

-

Industrial Fellowships Awarded to ten of the UK's most talented young researchers
-

-

-

-

UK Industry receives cash infusion from Prince Albert’s Royal Commission to tackle global challenges
-

Alumni
MoreThe Royal Commission for the Exhibition of 1851 has been awarding fellowships and scholarships since 1891. Previous holders of these prestigious awards include 13 Nobel Laureates and many more have gone on to become eminent in their field. Today the 1851 Alumni Network contains nearly 900 active members. Through events and online the Commission encourages cooperation and a crosspollination of ideas amongst its alumni and between alumni and current award holders.
Ernest Rutherford
1851 Award held 1895 – 1898
Nobel Prize in Chemistry 1908

